Projects Projects
Code, datasets, and research artefacts — from federated learning frameworks to proteomic resources used by thousands of researchers.
The sharing of sensitive patient data across institutions is a major bottleneck in cancer research. To address this, we developed a federated deep learning framework that enables collaborative training of cancer subtyping models without sharing raw data. By keeping data local and only sharing model updates, we can leverage the collective power of distributed datasets while preserving patient privacy. Our results demonstrate that federated models achieve performance comparable to centralized training, paving the way for large-scale, multi-institutional collaborations in precision oncology. This project culminated in the first application of federated learning to cancer proteomics, published in *Cancer Discovery*.
Federated LearningCancer SubtypingDeep Learning
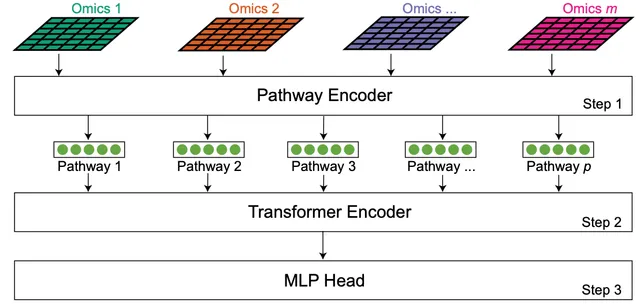
Multi-omic data analysis incorporating machine learning has the potential to significantly improve cancer diagnosis and prognosis. Traditional machine learning methods are usually limited to omic measurements, omitting existing domain knowledge, such as the biological networks that link molecular entities in various omic data types. Here we develop a Transformer-based explainable deep learning model, DeePathNet, which integrates cancer-specific pathway information into multi-omic data analysis. Using a variety of big datasets, including ProCan-DepMapSanger, CCLE, and TCGA, we demonstrate and validate that DeePathNet outperforms traditional methods for predicting drug response and classifying cancer type and subtype. Combining biomedical knowledge and state-of-the-art deep learning methods, DeePathNet enables biomarker discovery at the pathway level, maximizing the power of data-driven approaches to cancer research.
Deep LearningTransformerMulti-omics
Generative AI · Data Augmentation
High-quality multi-omic data is scarce and expensive to generate. To overcome this limitation, we developed MOSA (Multi-Omic Synthetic Augmentation), a generative AI model based on variational autoencoders. MOSA learns the underlying distribution of multi-omic data and generates realistic synthetic profiles that can be used to augment existing datasets. This approach increases the statistical power of downstream analyses, such as biomarker discovery and drug response prediction, and helps to address the issue of missing data in multi-omic integration. Our work, published in *Nature Communications*, demonstrates the potential of generative AI to enhance cancer data science.
Generative AIData AugmentationMulti-omics
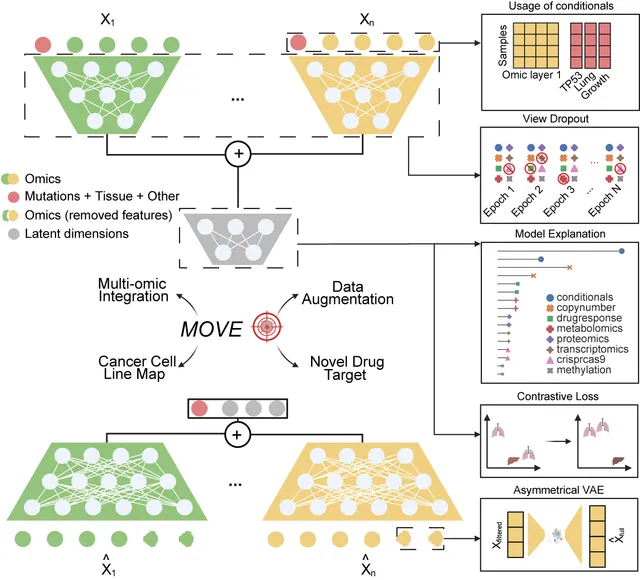
Integrative analysis of multi-omic datasets remains a challenge due to gaps and heterogeneity. We present a bespoke unsupervised deep learning model that generates synthetic multi-omic data for 1,523 cancer cell lines, completing the gaps and increasing the number of molecular and phenotypic profiles by 32.7%. Our model augments cellular measurements, improves cancer type clustering, and increases statistical power for cancer dependency biomarker discovery. Model explanation facilitates biomarker discovery and cancer target prioritization.
Deep learningMulti-view Variational AutoencoderMulti-omic integration
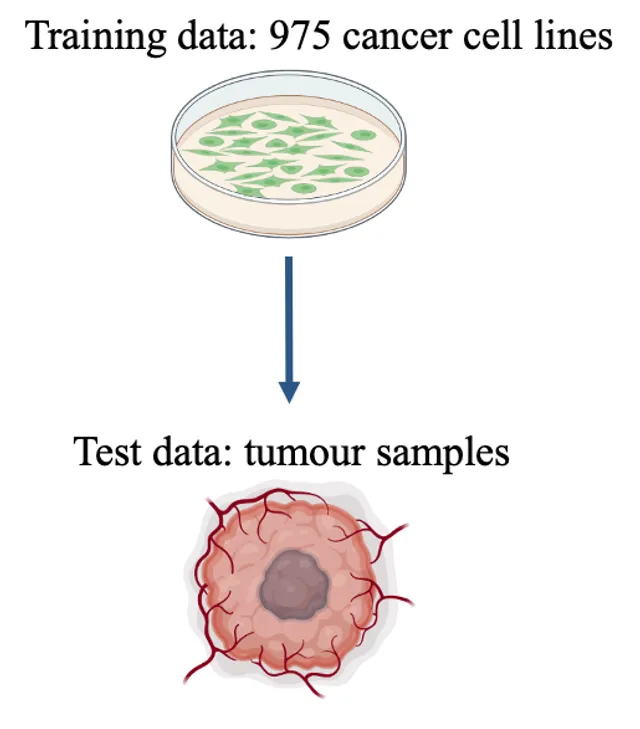
Cancer type is determined via assessment of tumour morphology, aided by immunohistochemical staining patterns. The development of machine learning (ML) models using histology slides has powered the image-based prediction of the site of origin in cancer of unknown primary (CUP). Here, we present an ML-based method to predict cancer type from a pan-cancer cohort consisting of 1,289 human tissue samples spanning 44 cancer types and 26 different tissues based on proteomic data. All samples were processed using data-independent acquisition mass spectrometry (DIA-MS). Two proteomic profiles from the pan-cancer cell line cohort were generated using two different sample preparation methods. These were normalized and merged by averaging the protein abundance, yielding a single training set (D1) with 975 cell lines and 9,688 proteins. Similarly, 1,277 tissue samples were processed by DIA-MS, quantifying 9,501 proteins. We trained a classifier using the cell lines (D1) as the baseline training set, and consecutively added 10% of D2 to D1 for online ML. We tested the baseline model and each subsequent new model on the test set T1. We observed a monotonic performance increase from 0.89 (baseline; Top-1 accuracy) to 0.97 (all D2 were used) when predicting the six cancer types. We observed an analogous trend when predicting the seven tissue types (from 0.64 to 0.84). Our proteomic-based ML model can predict cancer type and carcinoma tissue of origin in concordance with existing histopathological classification. It can also assign multiple probabilities to tumour type and tissue of origin, potentially enabling the classification of challenging pathology cases, such as CUP in future work. By adding tissue samples stepwise to the existing model, its predictive performance can be further enhanced. This reflects a real-world knowledge base that will continue to increase in predictive power with additional incremental proteomic data..
Machine learningCancer unknown primaryProteomics
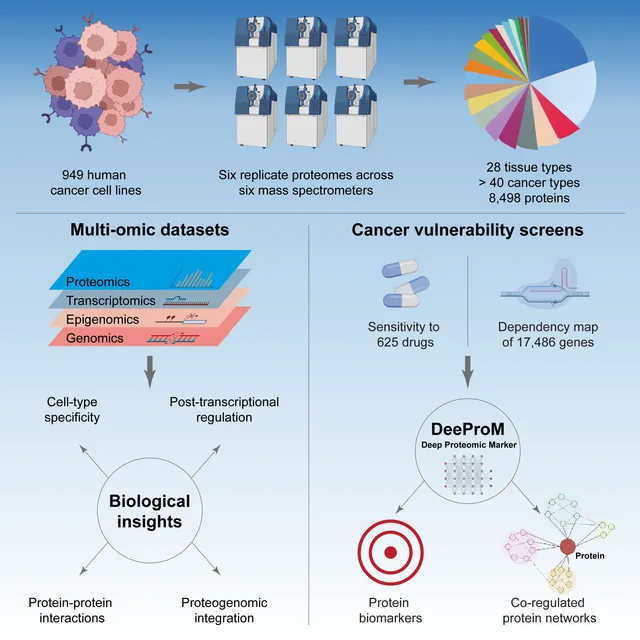
Proteomic data can reveal novel associations between genotype and phenotype, beyond what is apparent from genomics or transcriptomics alone. However, a lack of large proteomic datasets across a range of cancer types has limited our understanding of proteome network organisation and regulation. We produced a pan-cancer proteomic map derived from 949 human cancer cell lines. The map encompasses more than 40 cancer types derived from over 28 distinct human tissues. The samples were processed with a clinically-relevant workflow involving rapid and minimally complex sample preparation, quantifying 8,500 proteins. The raw proteomic data were acquired by data independent acquisition mass spectrometry (DIA-MS) at ProCan® in Australia. The processed data were analysed with a bespoke deep learning-based pipeline (DeeProM) that integrates multi-omics, CRISPR-Cas9 gene essentiality and drug sensitivity information produced at the Wellcome Sanger Institute. First, our findings reveal pervasive post-transcriptional modification and thousands of putative protein biomarkers of cancer vulnerabilities. Second, DeeProM statistics show that a fraction of the proteome can confer similar predictive power to the entire transcriptome. This has key implications for the clinical application of proteomics in drug response prediction. Third, we demonstrate that a random proportion of the identified proteins can provide robust predictions of cancer cell phenotypes, underpinning the concept of pervasive co-regulation of protein networks. This pan-cancer cell line proteomic map is a comprehensive resource that expands our understanding of cancer proteomes. These data reveal principles of cancer cell phenotypes, including genetic vulnerabilities and drug sensitivities, that are important for developing novel targeted anticancer therapies.
Deep learningMachine learningCancer
Deep learning · Natural language processing
The spread of misinformation can undermine confidence in vaccination. The capacity for social media to quickly and effectively spread information or misinformation is a pressing question for governments and global agencies. Tools such as deep learning, machine learning, sentiment analysis and text mining could provide solutions to help monitor the emergence of vaccine misinformation that underpin mitigation strategies and limit damage to vaccine confidence. You are invited to assist Assoc. Prof. Adam Dunn from the Faculty of Medicine and Health in designing a classifier to find vaccine sentiment in Twitter posts.
Deep learningNatural language processing
Deep learning · Computer vision
In ophthalmology, fundus screening is an economic and effective way to prevent blindness as early as possible that cased by diabetes, glaucoma, cataract, age-related macular degeneration (AMD) and many other causes. The figure below shows five simulated eye conditions when they have corresponding disease compared with normal vision.
Deep learningComputer vision
Single-cell RNA sequencing · Machine learning
I designed a new method which can be used by embryologists to determine the embryo differentiation propensity, given bulk RNA-seq data or qPCR data. The method is novel in using combined different single-cell datasets as a reference, then mapping the bulk RNA data to the lineage tree that is inferred from the single-cell data. This new method will greatly facilitate experiments where only bulk sequencing can be used.
Single-cell RNA sequencingMachine learningEmbryology
Deep learning · Natural language processing
KPMG and Melbourne Business School held a data science challenge with a topic in Natural Language Processing (NLP). The task was to explore complaints data from banks and analyse how NLP techniques can be leveraged. In this project, I explored and compared a number of NLP techniques, including tokenising, stemming, bag-of-words, word embedding etc, and designed a model to automatically classify complaint categories, so banks can use the model to increase workflow efficiency. The project won the 1st place with a $4000 prize.
Deep learningNatural language processing
Machine learning · Presto
During the 5 weeks internship at SEEK, together with another three group members and one mentor from SEEK, we developed a model to make prioritizations for the relevant jobs selected by existing algorithms. We started the project from the data collection stage and finished the project with a functional model which could lead to a potential 12.8% uplift on the click-through ratio with our estimation. The final dataset we focused contained over 400K observations with 1800+ features. We used both Presto and MS SQL server to extract the data needed. The final model was based on LightGBM, which works especially well on data with a large number of features. With a programming background, I did most of the technical work in the team, including AWS setup/troubleshooting, model tuning etc.
Machine learningPrestoAWS (Linux)




