Synthetic augmentation of cancer cell line multi-omic datasets using unsupervised deep learning
Integrative analysis of multi-omic datasets remains a challenge due to gaps and heterogeneity. We present a bespoke unsupervised deep learning model that generates synthetic multi-omic data for 1,523 cancer cell lines, completing the gaps and increasing the number of molecular and phenotypic profiles by 32.7%. Our model augments cellular measurements, improves cancer type clustering, and increases statistical power for cancer dependency biomarker discovery. Model explanation facilitates biomarker discovery and cancer target prioritization.
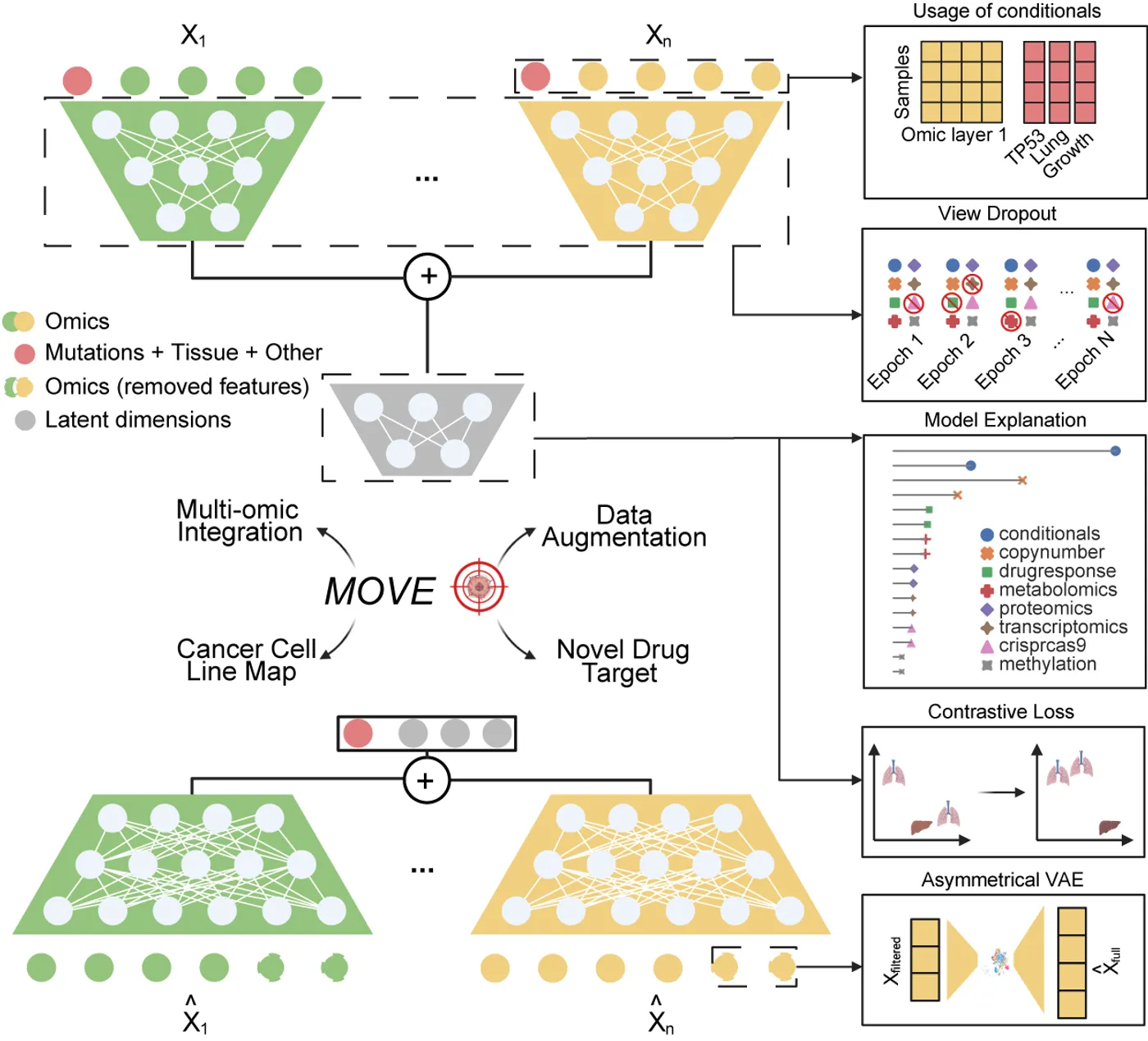
Integrative analysis of multi-omic datasets remains a challenge due to gaps and heterogeneity. We present a bespoke unsupervised deep learning model that generates synthetic multi-omic data for 1,523 cancer cell lines, completing the gaps and increasing the number of molecular and phenotypic profiles by 32.7%. Our model augments cellular measurements, improves cancer type clustering, and increases statistical power for cancer dependency biomarker discovery. Model explanation facilitates biomarker discovery and cancer target prioritization.